
Python in NeuroImaging
Find the community @
Stimuli Delivery
PsychoPyhttp://www.psychopy.orgPsychoPy is an easy, precise, platform-independent package for stimulus presentation. Suitable for psychophysics, neuroimaging, and all areas of psychology.∙ Huge variety of stimuli generated in real-time∙ Cross-platform – run the same script on Linux, Win or OS X∙ Flexible stimulus units (degrees, cm, or pixels)∙ Coder interface for those that like to program∙ Builder interface for those that don’t∙ Input from keyboard, mouse, joystick or button boxes∙ Multi-monitor support∙ Automated monitor calibration (supported photometers)
OpenSesamehttp://www.cogsci.nl/software/opensesameOpenSesame is a graphical experiment builder for the social sciences.∙ A comprehensive and intuitive graphical user interface∙ WYSIWYG drawing tools for creating visual stimuli∙ Cross-platform∙ Python scripting for complex tasks∙ A plug-in framework∙ Compatibility (through plug-ins) with commonly used devices: (e.g. Eyelink eye trackers, serial response boxes, Mantra object tracker)∙ Compatibility with popular Python libraries: PsychoPy, PyGame, PyOpenGL, etc.
Data I/O
NiBabelhttp://nipy.org/nibabelNibabel provides read and write access to some common medical and neuroimaging file formats, including: ANALYZE (plain, SPM99, SPM2), GIFTI, NIfTI1, MINC, as well as PAR/REC. NiBabel is the successor of PyNIfTI.The various image format classes give full or selective access to header (meta) information and access to the image data is made available via NumPy arrays.
Analysis
BrainVISAhttp://brainvisa.infoBrainVISA is an open-source, modular and customizable software platform built to host heterogeneous tools dedicated to neuroimaging research. It aims at helping researchers in developing new neuroimaging tools, sharing data and distributing their software.∙ Databasing capabilities∙ Massive computation facilities using Soma-workflow∙ Open environment, with many toolboxes∙ Specialized toolboxes for T1 MRI processing, sulci and gyri morphometry, diffusion imaging and fibers tracking, surfacic and structural analysis, 3D histology. . .∙ Links with other software like SPM, FSL, FreeSurfer, or CIVETD. Geffroy, D. Rivière, I. Denghien, N. Souedet, S. Laguitton, and Y. Cointepas. BrainVISA: a complete software platform for neuroimaging. In Python in Neuroscience workshop, Paris, Aug. 2011.
DiPyhttp://nipy.org/dipyDipy is an international FOSS project for diffusion magnetic resonance imaging analysis. Dipy is multiplatform and will run under any standard operating system such as Windows, Linux, Mac OS X. Some of our state-of-the-artapplications are:∙ Reconstruction algorithms e.g. GQI, DTI∙ Tractography generation algorithms e.g. EuDX∙ Intelligent downsampling of tracks∙ Ultra fast tractography clustering∙ Resampling datasets with anisotropic voxels to isotropic∙ Visualizing multiple brains simultaneously∙ Finding track correspondence between different brains∙ Warping tractographies into another (e.g. MNI) space∙ Support of various file formats e.g. Trackvis or NIfTI
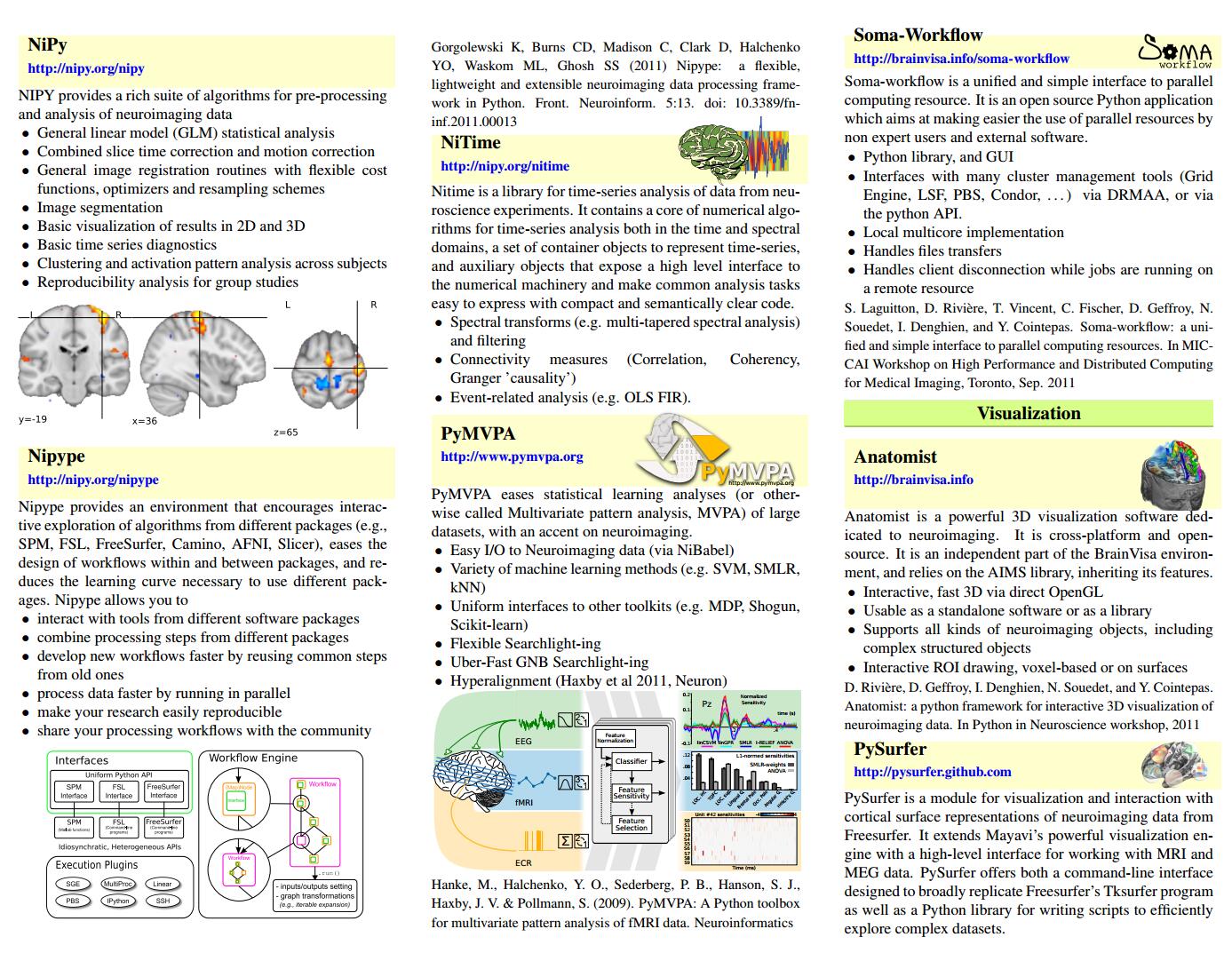
NiPyhttp://nipy.org/nipyNIPY provides a rich suite of algorithms for pre-processing and analysis of neuroimaging data∙ General linear model (GLM) statistical analysis∙ Combined slice time correction and motion correction∙ General image registration routines with flexible cost functions, optimizers and resampling schemes∙ Image segmentation∙ Basic visualization of results in 2D and 3D∙ Basic time series diagnostics∙ Clustering and activation pattern analysis across subjects∙ Reproducibility analysis for group studies
Nipypehttp://nipy.org/nipypeNipype provides an environment that encourages interactive exploration of algorithms from different packages (e.g., SPM, FSL, FreeSurfer, Camino, AFNI, Slicer),
eases the design of workflows within and between packages, and reduces the learning curve necessary to use different packages. Nipype allows you to∙ interact with tools from different software packages∙ combine processing steps from different packages∙ develop new workflows faster by reusing common steps from old ones∙ process data faster by running in parallel∙ make your research easily reproducible∙ share your processing workflows with the communityGorgolewski K, Burns CD, Madison C, Clark D, Halchenko YO, Waskom ML, Ghosh SS (2011) Nipype: a flexible, lightweight and extensible neuroimaging data processing framework in Python. Front. Neuroinform. 5:13. doi: 10.3389/fninf.2011.00013
NiTimehttp://nipy.org/nitimeNitime is a library for time-series analysis of data from neuroscience experiments. It contains a core of numerical algorithms for time-series analysis both in the time and spectral domains,a set of container objects to represent time-series, and auxiliary objects that expose a high level interface to the numerical machinery and make common analysis tasks easy to express with compact and semantically clear code.
∙ Spectral transforms (e.g. multi-tapered spectral analysis) and filtering∙ Connectivity measures (Correlation, Coherency, Granger ’causality’)∙ Event-related analysis (e.g. OLS FIR).
PyMVPAhttp://www.pymvpa.orgPyMVPA eases statistical learning analyses (or otherwise called Multivariate pattern analysis, MVPA) of large datasets, with an accent on neuroimaging.
∙ Easy I/O to Neuroimaging data (via NiBabel)∙ Variety of machine learning methods (e.g. SVM, SMLR, kNN)∙ Uniform interfaces to other toolkits (e.g. MDP, Shogun, Scikit-learn)∙ Flexible Searchlight-ing∙ Uber-Fast GNB Searchlight-ing∙ Hyperalignment (Haxby et al 2011, Neuron)Hanke, M., Halchenko, Y. O., Sederberg, P. B., Hanson, S. J., Haxby, J. V. & Pollmann, S. (2009). PyMVPA: A Python toolbox for multivariate pattern analysis of fMRI data. NeuroinformaticsHanke, M., Halchenko, Y. O., Sederberg, P. B., Hanson, S. J., Haxby, J. V. & Pollmann, S. (2009). PyMVPA: A Python toolbox for multivariate pattern analysis of fMRI data. NeuroinformaticsSoma-Workflowhttp://brainvisa.info/soma-workflowSoma-workflow is a unified and simple interface to parallel computing resource. It is an open source Python application which aims at making easier the use of parallel resources by non expert users and external software.∙ Python library, and GUI∙ Interfaces with many cluster management tools (Grid Engine, LSF, PBS, Condor, . . . ) via DRMAA, or via the python API.∙ Local multicore implementation∙ Handles files transfers∙ Handles client disconnection while jobs are running on a remote resource S. Laguitton, D. Rivière, T. Vincent, C. Fischer, D. Geffroy, N. Souedet, I. Denghien, and Y. Cointepas. Soma-workflow: a unified and simple interface to parallel computing resources. In MICCAI Workshop on High Performance and Distributed Computing for Medical Imaging, Toronto, Sep. 2011
Visualization
Anatomisthttp://brainvisa.infoAnatomist is a powerful 3D visualization software dedicated to neuroimaging. It is cross-platform and opensource. It is an independent part of the BrainVisa environment, and relies on the AIMS library, inheriting its features.∙ Interactive, fast 3D via direct OpenGL∙ Usable as a standalone software or as a library∙ Supports all kinds of neuroimaging objects, including complex structured objects∙ Interactive ROI drawing, voxel-based or on surfaces D. Rivière, D. Geffroy, I. Denghien, N. Souedet, and Y. Cointepas. Anatomist: a python framework for interactive 3D visualization of neuroimaging data. In Python in Neuroscience workshop, 2011
PySurferhttp://pysurfer.github.comPySurfer is a module for visualization and interaction with cortical surface representations of neuroimaging data from Freesurfer. It extends Mayavi’s powerful visualization engine with a high-level interface for working with MRI and MEG data. PySurfer offers both a command-line interface designed to broadly replicate Freesurfer’s Tksurfer program as well as a Python library for writing scripts to efficiently explore complex datasets.